SemLab and Catharina Hospital develop RAG-tool for medical protocols
According to internal research, nurses at Catharina Hospital spend around 3 hours per department per day
consulting medical documentation such as protocols and guidelines. While some of this time is spent reading and
processing the relevant information, valuable time is lost searching through various sources and collecting
fragmented data, instead of using it on patient care. Also, because nurses are under immense time pressure,
scattered documentation might lead to it not being thoroughly consulted, or at all, which could degrade the
quality and consistency of care.
WIthin the context of the MedGPT-project, SemLab and Catharina Hospital are working together to combat this
issue and develop a tool to decrease the time nurses spent on information gathering.
Large Language Models
In recent years Large Language Models (LLM’s) have become increasingly popular aides for (among other uses)
question answering and information retrieval. More and more people rely on them for both personal and
professional tasks. However, while undeniably powerful, out-of-the-box LLM’s are often designed with general
purpose in mind and lack domain- and task-specific capabilities.
For medical professionals at Catharina Hospital it is imperative only selected, trusted sources are used when
looking up information. Which sources are used also needs to be transparent so answers are verifiable and
relevant documents is easily accessible. Lastly, a very high degree of certainty that answers are correct is
required as any wrong or missing information could have dire consequences. So models need to be overly cautious
in whether they can safely answer a query, which contrasts with the directive of most LLM’s that aim to always
have an answer for a user.
Retrieval Augmented Generation
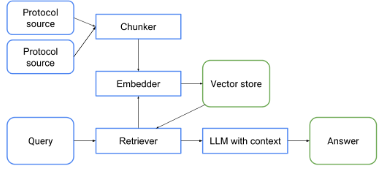
To create a tool that fullfills these requirements, SemLab is developing a Retrieval Augmented Generation
(RAG)-system specifically tailored for Catharina Hospital. A RAG-system is still a system that provides an
answer to a question, but does so via a multi-step process. First, all relevant sources, provided by Catharina
Hospital, are split into coherent chunks and stored in a vector database. When a user enters a query, a
retriever-model gets chunks from this database that contain relevant information for this query. These chunks
are then provided to a LLM as context based on which it needs to answer the query. Additional guardrails will be
implemented around all steps to check that there are relevant sources available, the LLM indeed did base its
answer solely on these, and there is a high degree of reliability and certainty for the answer.
Development
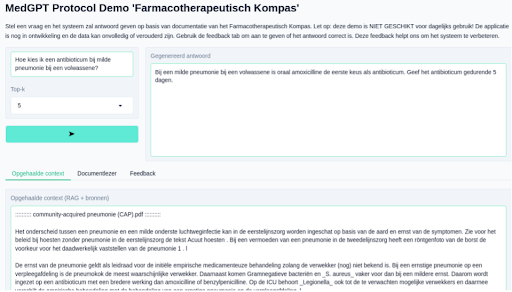
Initial development by SemLab is based on publicly available protocols from
https://www.farmacotherapeutischkompas.nl/ and synthetically generated queries. The aim is to soon deploy a
first prototype on-premise at Catharina Hospital. The goal of this rapid prototyping is for medical
professionals to interact, test and experiment with the tool as early as possible, and on actual data from
Catharina Hospital, as their feedback is essential to steer improvements and next steps. This development cycle
combines the technical proficiency from SemLab and medical expertise and day-to-day use from Catharina Hospital
to build a reliable and efficient tool for querying medical protocols.
Posted by SemLab on 17 april 2026